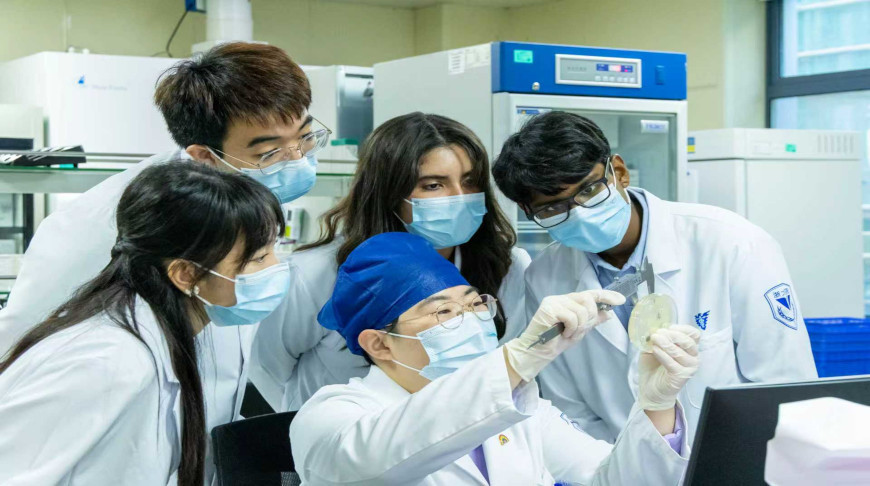
Photo: ZJUISM / iStock
BEIJING, 25 March (BelTA - China Daily) - A multidisciplinary team
of researchers from the Hangzhou-based Zhejiang University have made a
major breakthrough in membrane proteins that could pave the way for
tackling genetic afflictions like Parkinson's disease.
The study, published in Nature last month, shows the researchers have successfully engineered a set of artificial proteins that are able to regulate the functions of G protein-coupled receptors, or GPCRs.
More than 30 percent of approved drugs worldwide target these receptors, said Zhang Yan, vice-dean of Zhejiang University's School of Medicine and one of the leading researchers of the study.
Abnormal genetic expression and mutations in the receptors can impair these switches and therefore disrupt their signaling functions. In fact, hundreds of clinical diseases, including Parkinson's, obesity and hypercalcemia (high level of calcium in the blood), are found to be caused by such mutations.
For patients, these structural dysfunctions often translate into long-term and chronic burdens, as conventional drugs designed to target the receptors' switches are generally unable to execute repairs.
That is where artificial intelligence comes to the researchers' aid, as the development of AI-driven protein design in recent years, particularly generative models for de novo design, has provided tools that generate proteins with unprecedented speed and accuracy.
The goal of de novo design — making new proteins from scratch - is to design proteins that do not exist in nature, without relying on natural structures or sequences, according to Zhang Min, another member of the research team from Zhejiang University's College of Computer Science and Technology.
Compared to modifying or improving existing proteins, this task presents greater challenges for researchers and algorithms in understanding proteins.
"We need to systematically deconstruct a seemingly simple requirement and transform it into functional modules that AI can implement step by step," she said.
However, the real hurdle lies in creating the right modulators to bind the receptors at the right places and achieve the desired effects.
To address that challenge, the team developed an AI-guided probe, through which the structures of the targeted receptors can be thoroughly profiled and potential binding and regulatory sites found. Then, using an approach known as "structural prompts" - not dissimilar to input fed to Deep-Seek or ChatGPT, but for protein structures - they generated these modulators.
More significantly for the research team, their findings can now serve as a platform for similar research.
The study, published in Nature last month, shows the researchers have successfully engineered a set of artificial proteins that are able to regulate the functions of G protein-coupled receptors, or GPCRs.
More than 30 percent of approved drugs worldwide target these receptors, said Zhang Yan, vice-dean of Zhejiang University's School of Medicine and one of the leading researchers of the study.
Abnormal genetic expression and mutations in the receptors can impair these switches and therefore disrupt their signaling functions. In fact, hundreds of clinical diseases, including Parkinson's, obesity and hypercalcemia (high level of calcium in the blood), are found to be caused by such mutations.
For patients, these structural dysfunctions often translate into long-term and chronic burdens, as conventional drugs designed to target the receptors' switches are generally unable to execute repairs.
That is where artificial intelligence comes to the researchers' aid, as the development of AI-driven protein design in recent years, particularly generative models for de novo design, has provided tools that generate proteins with unprecedented speed and accuracy.
The goal of de novo design — making new proteins from scratch - is to design proteins that do not exist in nature, without relying on natural structures or sequences, according to Zhang Min, another member of the research team from Zhejiang University's College of Computer Science and Technology.
Compared to modifying or improving existing proteins, this task presents greater challenges for researchers and algorithms in understanding proteins.
"We need to systematically deconstruct a seemingly simple requirement and transform it into functional modules that AI can implement step by step," she said.
However, the real hurdle lies in creating the right modulators to bind the receptors at the right places and achieve the desired effects.
To address that challenge, the team developed an AI-guided probe, through which the structures of the targeted receptors can be thoroughly profiled and potential binding and regulatory sites found. Then, using an approach known as "structural prompts" - not dissimilar to input fed to Deep-Seek or ChatGPT, but for protein structures - they generated these modulators.
More significantly for the research team, their findings can now serve as a platform for similar research.